Our lab utilizes a protein engineering tool know as expanded genetic code technology. We have applied such technique to interrogate an array of nucleic acid binding proteins to better study RNA/DNA-protein interactions. This technology has allowed us to design tools to further understand the role of these proteins in context of viral infections.
Genetic code expansion technologies utilize unnatural amino acids (UAAs) beyond the 20 known canonical amino acids. Canonical tRNA synthetase/ tRNA pairs are evolved in lab settings to introduce and recognize the UAA of interest in response to a rare codon. This technology is known to be bioorthogonal, with minimal to little interruptions to the system’s natural processes.
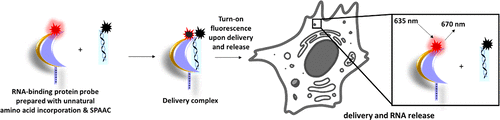
Our lab has utilized this technology to further engineer p19 for the detection of its small non-coding RNA ligands. This was done by incorporating Azidophenylalanine, site specifically. Through biorthogonal click chemistry, we were able to engineer a Förster resonance energy transfer (FRET) probe for the detection of binding and delivery of RNA cargo into cells.
Genetic code expansion technologies utilize unnatural amino acids (UAAs) beyond the 20 known canonical amino acids. Canonical tRNA synthetase/ tRNA pairs are evolved in lab settings to introduce and recognize the UAA of interest in response to a rare codon. This technology is known to be bioorthogonal, with minimal to little interruptions to the system’s natural processes.
Our lab has utilized this technology to further engineer p19 for the detection of its small non-coding RNA ligands. This was done by incorporating Azidophenylalanine, site specifically. Through biorthogonal click chemistry, we were able to engineer a Förster resonance energy transfer (FRET) probe for the detection of binding and delivery of RNA cargo into cells.
We have, additionally, taken advantage of this technology to engineer super suppressors of the RNA silencing pathway. This was done by the site-specific incorporation of a photoactive unnatural amino acid into the p19 dimer, which led to cross-linking between the protein and its targets. The engineered protein demonstrated the possibility of using such technology to irreversibly sequester small non-coding RNAs through covalent cross-linking.
Similar to the p19-FRET probe, we engineered a hepatitis C virus nonstructural protein 3 helicase (NS3h) based-FRET probe to study its helicase function. We utilized genetic code expansion technology to engineer a protein that allows for smFRET experiments. The engineered probe allowed us to monitor the dynamics of enzyme translocation during unwinding in real-time.
Select research articles:
Ahmed, N.; Ahmed, N.; Bilodeau, D. A.; Pezacki, J. P. "An unnatural enzyme with endonuclease activity towards small non-coding RNAs" Nat. Comm. 2023, 14, 1, 3777. DOI: 10.1038/s41467-023-39105-0
Ablenas, C., Gidi, Y., Powdrill, M., Ahmed, N., Shaw T., Mesko, M., Gotte, M., Cosa, G., Pezacki, J.P. “Hepatitis C virus helicase binding and activity monitored through site-specific labeling using an expanded genetic code.” ACS Infect. Dis. 2019, 5, 2118−2126
Ahmed, N.; Foss, D.V.; Powdrill, M.H.; Pezacki JP “Site-specific crosslinking of a p19 viral suppressor of RNA silencing protein and its RNA targets using an expanded genetic code” Biochemistry 2019, 58, 3520-3526
Ahmed N, De Graaf JF, Ahmed N, Foss DV, Delcorde J, Schultz PG, Pezacki JP "Visualization of the delivery and release of small RNAs using genetic code expansion and unnatural RNA-binding proteins " ACS Bioconjug. Chem. 2018, 29(12), 3982-3986.
Similar to the p19-FRET probe, we engineered a hepatitis C virus nonstructural protein 3 helicase (NS3h) based-FRET probe to study its helicase function. We utilized genetic code expansion technology to engineer a protein that allows for smFRET experiments. The engineered probe allowed us to monitor the dynamics of enzyme translocation during unwinding in real-time.
Select research articles:
Ahmed, N.; Ahmed, N.; Bilodeau, D. A.; Pezacki, J. P. "An unnatural enzyme with endonuclease activity towards small non-coding RNAs" Nat. Comm. 2023, 14, 1, 3777. DOI: 10.1038/s41467-023-39105-0
Ablenas, C., Gidi, Y., Powdrill, M., Ahmed, N., Shaw T., Mesko, M., Gotte, M., Cosa, G., Pezacki, J.P. “Hepatitis C virus helicase binding and activity monitored through site-specific labeling using an expanded genetic code.” ACS Infect. Dis. 2019, 5, 2118−2126
Ahmed, N.; Foss, D.V.; Powdrill, M.H.; Pezacki JP “Site-specific crosslinking of a p19 viral suppressor of RNA silencing protein and its RNA targets using an expanded genetic code” Biochemistry 2019, 58, 3520-3526
Ahmed N, De Graaf JF, Ahmed N, Foss DV, Delcorde J, Schultz PG, Pezacki JP "Visualization of the delivery and release of small RNAs using genetic code expansion and unnatural RNA-binding proteins " ACS Bioconjug. Chem. 2018, 29(12), 3982-3986.