Computational Enzyme Design
Our lab develops computational methods for the design of efficient artificial enzymes, while also investigating the fundamental principles of enzyme catalysis. Using both experimental and computational approaches, we explore how conformational dynamics shape catalytic cycles and use these insights to guide new design strategies. Our long-term goal is to create a unified platform for enzyme design, capable of producing efficient biocatalysts for virtually any reaction, while training the next generation of scientists in cutting-edge protein engineering.
Latest Results
Hunt et al. (2025) Journal of the American Chemical Society.
Distal Mutations in a Designed Retro-Aldolase Alter Loop Dynamics to Shift and Accelerate the Rate-Limiting Step.
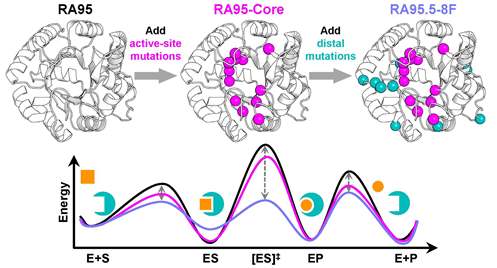
Amino-acid residues distant from an enzyme's active site are known to influence catalysis, but their mechanistic contributions to the catalytic cycle remain poorly understood. Here, we investigate the structural, functional, and mechanistic impacts of distal and active-site mutations discovered through directed evolution of the computationally designed retro-aldolase RA95. Active-site mutations improve catalytic efficiency by 3,600-fold, while distal mutations alone offer no improvement. When combined with active-site mutations, distal mutations further increase efficiency by 6-fold, demonstrating an epistatic effect. X-ray crystallography and molecular dynamics simulations reveal that distal mutations promote active site opening by altering loop dynamics. Kinetic solvent viscosity effects and electric field calculations show that distal mutations accelerate the chemical transformation by 100-fold, shifting the rate-limiting step to product release, which is further accelerated by the increased opening of the active site. These findings establish the critical role of distal residues in shaping the active-site environment and facilitating the structural dynamics essential for efficient progression through the catalytic cycle, offering valuable insights for enzyme design.
Read the article here.
Previous results can be found here.
Research Funding
We gratefully acknowledge support from the following agencies:

Keywords
Protein Engineering, Computational Protein Design, Enzyme Design, Enzymes, Biocatalysis, Fluorescent Proteins, Protein Dynamics, Molecular Modeling, Protein Science, Biological Chemistry
Contact info
Roberto Chica, Ph. D. (613) 562-5800 x 1988 |
 |
|---|